The goal of this class is to help grad students, postdocs, or faculty who have a background in basic statistics and a familiarity with some other statistics package (JMP, SYSTAT, SAS, SPSS) to become comfortable with the R Project as a platform for statistical analyses. It is not meant as a course in statistics, nor does it cover more than a small portion of what is available in R. At the end of the course you should be comfortable managing data in R, making graphs, performing an array of different basic statistical analyses, and be able to use the extensive resources available on line and in books to learn to do just about anything statistical you'd like to in R.
The goal of this class is to help grad students, postdocs, or faculty who have a background in basic statistics and a familiarity with some other statistics package (JMP, SYSTAT, SAS, SPSS) to become comfortable with the R Project as a platform for statistical analyses. It is not meant as a course in statistics, nor does it cover more than a small portion of what is available in R. At the end of the course you should be comfortable managing data in R, making graphs, performing an array of different basic statistical analyses, and be able to use the extensive resources available on line and in books to learn to do just about anything statistical you'd like to in R.
The course is offered periodically, when enough UCSC grad students request it, but you can take the course on your own! The class is primarily based on the handouts below, and videos of the 2015 version lectures are available for UCSC students.
In 2016 I offered a 2-day version of the Transition to R class, covering just the basics to get people up and running in R. Here are the Documents from that class. ENJOY!
RBeginners_A simple analysis.PDF
somedata.csv (Google sheet: File>Download>as CSV file)
IMPORTANT NOTE ON LINKS TO DATA. The website has moved since this course was last offered, so the URLs to datasets in the handouts are broken. Please note, however, that all the datasets are still available, below. You can download them, or get their current URLs by right-clicking on the links, and then modifying the links in the Rcode. I will be updating this soon, but in the meantime...
Syllabus, class notes, and class videos
ENVS291 Transition To R Syllabus W2015.pdf
Here are the detailed handouts, and in-class videos of the lectures and conversations from the 2015 version of ENVS291 Transition to R. Use at your own risk. All handout and video contents are the 2015 copyright of Gregory S. Gilbert, University of California, Santa Cruz
Class1_GettingStarted.pdf LectureVideo
Installing R; R windows; loading libraries; R Types, structures, and objects; R terminology; basic R notation; common operators; exploring R objects; referencing inside data frames, vectors, and matrices; importing data from a spreadsheet
Class2_Handling Data.PDF LectureVideo
Creating, editing, reading, and exporting dataframes; sorting, subseetting, combining data; importing and exporting data; handling dates
Class3_Summarizing Data.PDF LectureVideo
Summarizing continuous data; functions in R; Creating customized complex functions; Summarize continuous data in groups; Apply functions across rows or columns; by, aggregate, tapply, apply, sapply, lapply
Classes 4 and 5 Regression, ANOVA, tTest.PDF LectureVideo4 LectureVideo5
Fitting models; Extractor functions; linear regression using lm; ANOVA using aov or lm; ttest and rank tests; factorial, blocked, and split-plot designs; ANCOVA; Homogeneity of variance; Type I and III sums of squares; predict for fitted lines and confidence intervals; stepwise selection
Class6_BasicPlottingTools.PDF LectureVideo
Base plotting tools; plot, boxplot, hist; plot overlays; par function to control plot attributes; exporting graphs
Class7 Functions and Loops.PDF LectureVideo
Custom functions; Basics of programming; Conditional statements; Loops; Writing scripts in R
Class8 GLMs and MixedModels.PDF LectureVideo
Generalized Linear models (glm); error structure, linear predictors, link functions; Logistic regression; Poisson regression; Survival Analysis; Mixed Models
Class9 Advanced Graphics with ggplot.PDF LectureVideo
Graphs with ggplot2; data, aesthetic attributes, mapping, geoms, stats, facets, layers, viewports, themes, etc.
Class10a Vegan Community Ecology Analysis.PDF no class video
Basics of Vegan package for analysis of community structure and diversity data. Measures of diversity; NMDS NOT UPDATED SINCE 2013; SOME PARTS ARE OUT OF DATE.
Class10b Picate and Phylomatic Phylogenetic Ecology tools.PDF no lecture video
Very brief introduction to tools for phylogenetic community ecology and trait analysis. NOT UPDATED SINCE 2013 SOME THINGS ARE OUT OF DATE! Newick tree format; making phylogenetic trees from phylomatic; phylogenetic diversity measures
FERP data used

Use the color guide at http://html-color-codes.info to get 6-character color codes, or RGB codes. Then plot(y~x, col="#8E2BC3") or mycol1<-rbg(142/255,43/255,195/255,.7); plot(y~x, pch=19,col=mycol1)
These are the data used for all the scatterplot examples. Copy and paste these into the R console, and hit return. Then copy and paste the code from any of the graphs to reproduce the graphs.
x<-c(1,2,3,4,5,6,7,8)
y1<-c(2,4,5,7,8,7,9,10)
y2<-c(1,3,2,4,6,5,7,7)
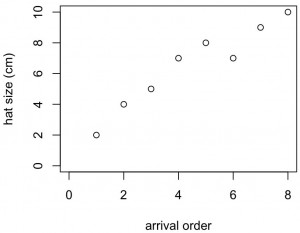 plot(x,y1,xlab="arrival order",ylab="hat size (cm)", ylim=c(0,10),xlim=c(0,8))
plot(x,y1,xlab="arrival order",ylab="hat size (cm)", ylim=c(0,10),xlim=c(0,8))
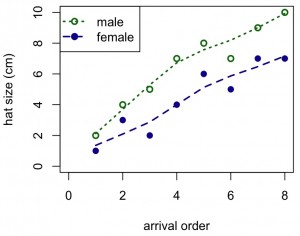 #add smooth lowess curves to each set of points in the scatterplot
#add smooth lowess curves to each set of points in the scatterplot
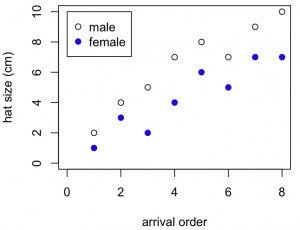
plot(x,y1,xlab="arrival order",ylab="hat size (cm)",ylim=c(0,10),xlim=c(0,8),col="dark green",pch=1,lwd=2)
lines(lowess(x,y1),lwd=2,lty=3,col="dark green")
points(x,y2,pch=19,col="dark blue")
lines(lowess(x,y2),lwd=2,lty=2,col="dark blue")
legend("topleft",c("male","female"),lty=c(3,2),pch=c(1,19),col=c("dark green
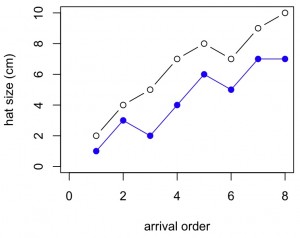
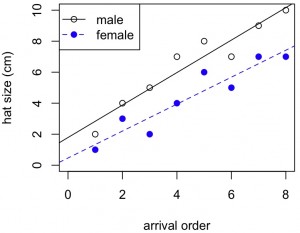 #Use abline to add linear regression lines to each set of points in the scatterplot
#Use abline to add linear regression lines to each set of points in the scatterplot
plot(x,y1,xlab="arrival order",ylab="hat size (cm)",ylim=c(0,10),xlim=c(0,8),col="black",pch=1,lwd=1)
abline(lm(y1~x),lwd=1,lty=1,col="black")
points(x,y2,pch=19,col="blue")
abline(lm(y2~x),lwd=1,lty=2,col="blue")
legend("topleft",c("male","female"),lty=c(1,2),pch=c(1,19),col=c("black","blue"),lwd=c(1,1))
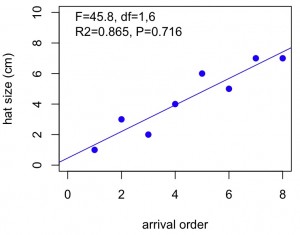 #get the relevant statistics for the regression line, then put on the graph as text
#get the relevant statistics for the regression line, then put on the graph as text
a<-summary(lm(y2~x)) #this puts summary stats of the linear regression of y2 on x into list a
R2<-signif(a$adj.r.squared,3) #adjusted R squared
F<-signif(a$fstatistic[1],3) #F statistic
ndf<-signif(a$fstatistic[2],1) #degrees of freedom numerator
ddf<- signif(a$fstatistic[3],1) #degress of freedom denominator
P<-signif(a$coefficients[2,4],4) #P value for significant slope
plot(x,y2,xlab="arrival order",ylab="hat size (cm)",ylim=c(0,10),xlim=c(0,8),col="blue",pch=19,lwd=1)
abline(lm(y2~x),lwd=1,lty=1,col="blue") #puts in the regression line
text(0,9,paste("F=",F,", df=",ndf,",",ddf,"n","R^2=",R2,", P=",P,sep=""),pos=4) #adds the statistics